・Realizing precision medicine for atrial fibrillation utilizing next generation sequencing and machine learning technologies
Susceptibility to atrial fibrillation (AF) has been reported to be determined not only by well-known risk factors such as hypertension, diabetes mellitus, and structural heart disease, but also by genetic variants. Recent advances in technology have led to the genotyping of single nucleotide polymorphisms (SNPs) across the genome, and genome-wide association studies (GWAS), which statistically examine the frequency of each SNP and its association with disease and traits, have increasingly become popular. As multi-ancestry genomic data with a large sample size is currently available, recent GWAS have reported a substantial number of loci associated with AF. However, the vast majority of GWAS significant loci remains unclear about their functionalities, and the genetic architecture of AF is not comprehensively understood.
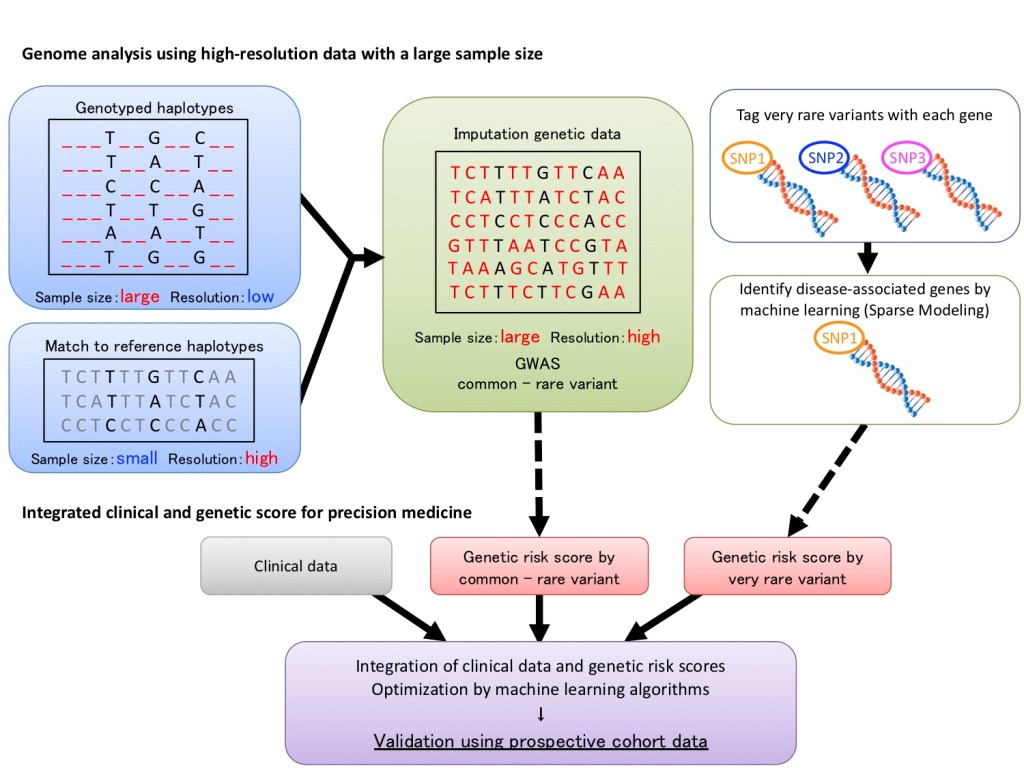
In this project, we aim to thoroughly elucidate pathophysiological mechanism underlying AF. We generated high-resolution genomic data, including common variants and rare variants, with a sufficient number of samples by imputation method on the genomic data of approximately 200,000 Japanese patients in BioBank Japan. In addition, for very rare variants that have very low frequencies but are inferred to have a large effect size, we will apply machine learning algorithms to identify the rare variants and perform comprehensive GWAS on a wide range of genetic variants to elucidate more detailed pathological mechanisms of AF.
Furthermore, in order to apply the genomic information to clinical practice, we aim to create a novel genetic risk score combining the obtained GWAS results with clinical data. The polygenic risk score has been shown to predict AF occurrence as well as adverse outcomes such as stroke. By sharing and integrating the information on multi-ancestry data with our collaborators, we will optimize genetic risk scores by machine learning algorithms and evaluate their performance. The best performing genetic risk score is validated using prospective cohort data and is applied to risk stratification for disease outcomes. Thus, appropriate treatment tailored to individual pathologies and disease risk can contribute to realization of next-generation precision medicine.